Introduction
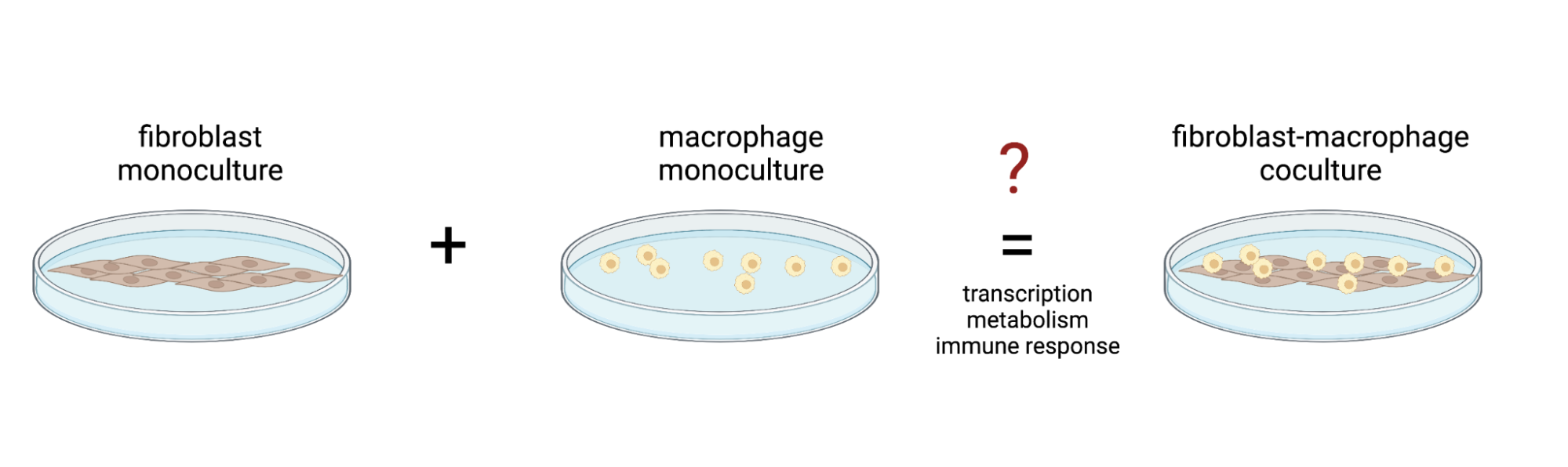
Cells live together in complex, interacting ecosystems. When we bring them into the lab, we strip them of these circumstances. We isolate, confine, and silo life. That’s why this summer, as a New Science fellow, I probed the mechanisms that allow cells to self-organize into higher order systems. I want to understand how cells change when they are grown in close proximity to other types of cells (Figure 1)
I’m fascinated by how complex, adaptive systems work, and how they fail. Most of my life has been spent in labs that study longevity and aging. Prior work studied the changes in methylation associated with aging, and it’s there that I tangentially learned of the strong association between methylation and cell types and also learned that cell compositions shift — the very identity of cells change — in both aging and disease.
To find a framework to compress these observations, I turned to theoretical biology, especially Leo Buss’ book on the evolution of multicellularity and Robert Rosen’s writings on ‘anticipatory systems.’ Rosen notes that it’s possible for a system to fail in the coordination of its parts, in the system’s efforts to anticipate their environment and themselves, rather than a failure in the parts themselves. Other scientists, like Victor de Lorenzo, have opined on “The Selfish Metabolism,” the notion that metabolism could play a critical role in driving the same phenotypes that genetics and epigenetics aim to control.
These ideas made me wonder: How do the local dynamics and coordination of individual cells lead to complex, higher order phenotypes?
I designed and began experiments to probe how individual cell behaviors contribute to the functionality of higher-level systems. Specifically, I co-cultured cells together and then quantified the shifts in metabolisms and gene expression that follow. For more thoughts on cell-cell cooperation and higher-order behaviors, see my essay for New Science.
Project Overview
During the summer, I focused on gaining experience in the various techniques I’d need to develop and build the necessary datasets to better understand cell-cell coordination from first principles. My work began with computational experiments, using available datasets, and then I learned to co-culture fibroblasts and macrophages as an experimental framework to ask deeper questions. Liquid Chromatography - Mass Spectrometry (LCMS) experiments aimed to provide an initial look into the metabolic aspects of monoculture versus coculture. Synthetic myddosomes, in the future, may be used to perturb intercellular signaling and metabolic cascades.
These milestones are broken into 4 discrete modules:
- 1. Computational exploration of metabolism.
- 2. Co-culturing fibroblasts and macrophages.
- 3. Pilot LCMS experiments.
- 4. Creating synthetic myddosomes.
Project Modules
Module 1: Computational Forays
My first step in this project was to systematically understand the ‘normal’ levels for various metabolites in different cells. Then, when these cells are co-cultured, we have a baseline to make comparisons about which specific metabolites change.
To start my project, I used an existing gene expression dataset that was collected by Xu Zhou. It contains data on the relative levels of different genes in monocultured fibroblasts and macrophages. I investigated whether genes that are highly expressed in one cell type, but lowly expressed in another, represented a candidate of interaction; a division of metabolic labor, if you will.
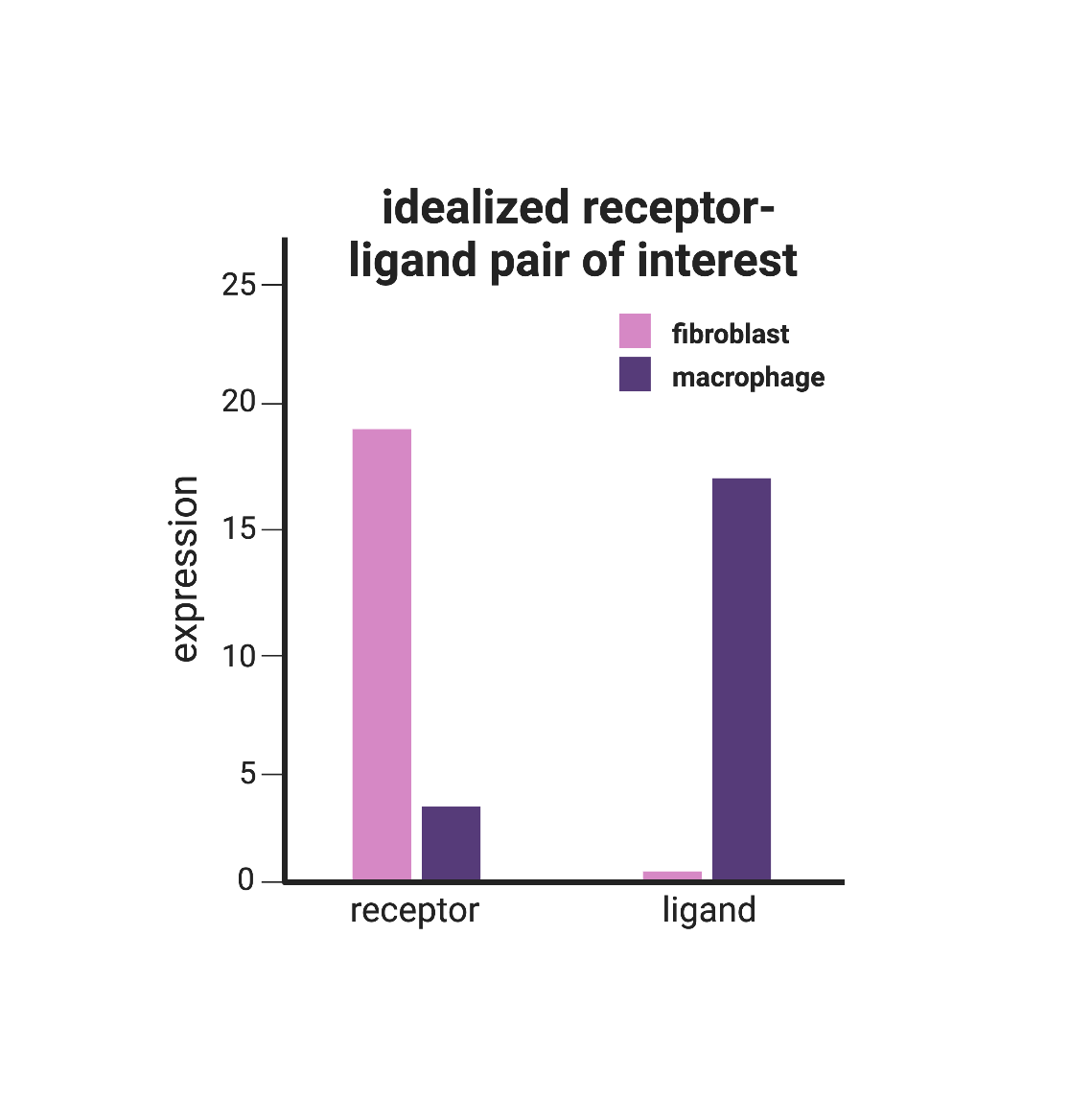
To begin searching for intercellular dependencies, I started with a list of ligand-receptor pairs, and calculated a log2diff value to score how differentially expressed a gene was between the cell types (Figure 2). This score is calculated using the equation:
where MEF and BMDM are fibroblasts and macrophages, respectively. This calculation was done twice within each row in the dataset; once for the ligand gene (log2diff.x) and once for the receptor gene (log2diff.y). The resulting list was then ordered by the ligand-receptor pairs that had the most discrepancy between gene expression values by cell type, via the calculation:
I ended up with a list of candidate ligand-receptor pairs (table in git repository; see the Appendix), and it was encouraging that the pairs seen in Xu Zhou’s 2-cell circuit paper were found among the top pairs (e.g. the genes CSF1, PDGFb).
I next repeated this analysis using a reference list of enzyme-enzyme pairs (KEGG), applying the same equation to score the pairs (table in git repository; see Appendix). To better visualize the recurrence of some enzymes found in the list, I graphed the top 30 pairs (Figure 3):
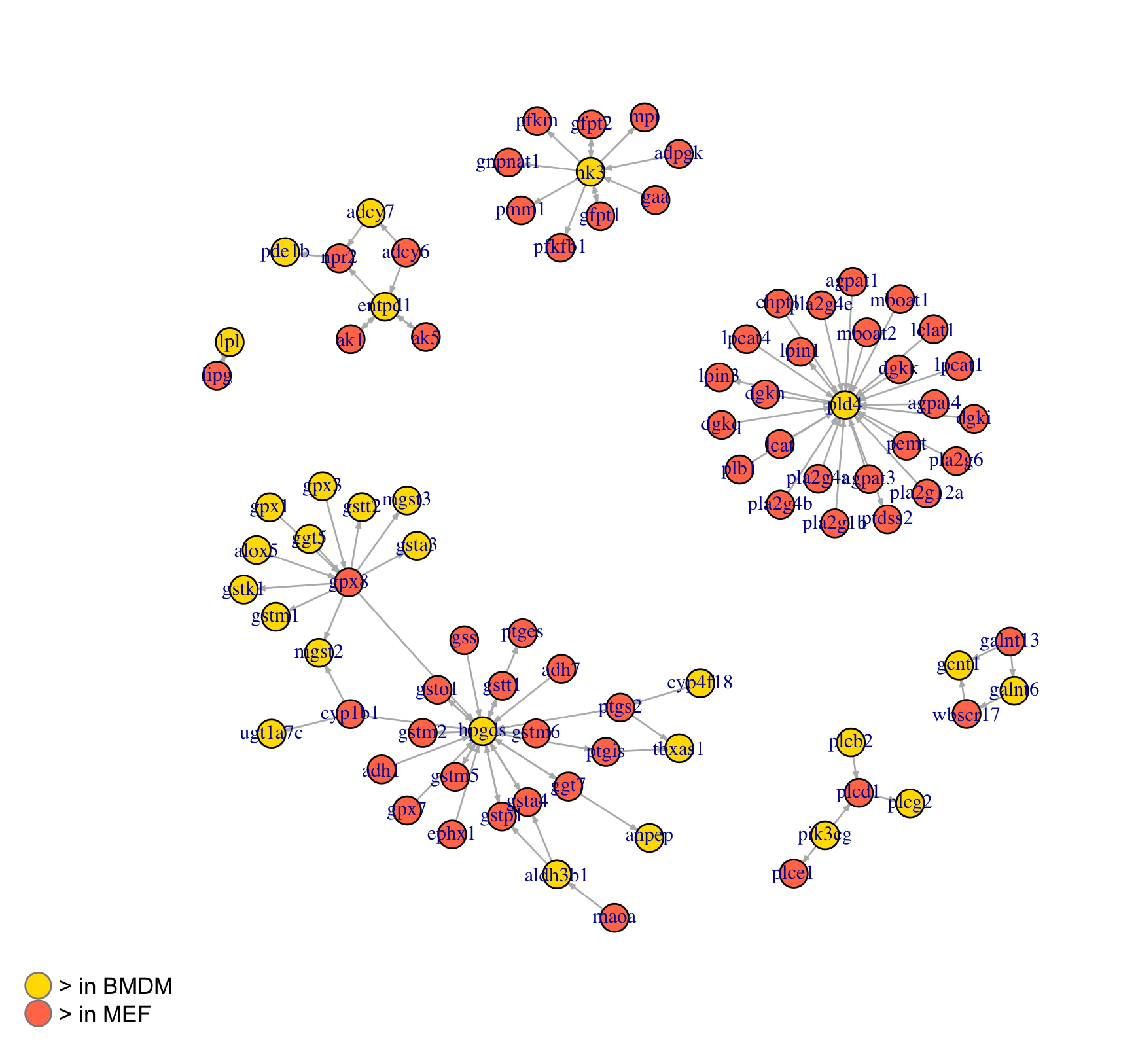
For this analysis, I had less intuition and references to determine whether my results were biologically significant. And there were some unanswered considerations. The ‘hub’ genes, for instance, are partially an artifact of having high expression values. And genes with high expression values will generally have higher pair differential scores than genes with lower gene expression. I’m unsure whether this obscures the rankings that I’m interested in. And even if these genes are strongly expressed, they need to be accessible to the other cell type (i.e. transported), and this aspect can’t be clarified using this set of analyses.
As another approach to probe at the biological significance of this analysis, I selected the enzyme pairs relevant to oxidative phosphorylation and glycolysis pathways and repeated my analysis. Specifically, I hypothesized that all the gene expressions in these pathways must be significant (because cells must be able to do at least one to survive, right?).
Surprisingly, I found many of them to be incomplete — some gene expressions had zero value. I don’t fully understand how to explain this, but it provokes the question: what is necessary for a complete cell circuit? I wonder if there might be an answer from the field of database querying, where there is invariance in how you transform a database from one into another; perhaps there are multiple paths to generate an essential end product? Or gene expression doesn’t correspond to the amount of gene product there is. But these were not followed up on much because I then had the opportunity to do benchwork.
Module 2: Coculture of fibroblasts and macrophages
To learn how to generate my own datasets of interest, I started experimental work by relearning tissue culture and aseptic techniques. I was also introduced to the fibroblast/macrophage experimental system first developed by Xu Zhou. These cells, when grown in co-culture, offer a relatively simple model system that exhibits emergent properties — the cells regulate their population ratios and exhibit different immune responses in coculture vs monocultures.
After analyzing Xu’s dataset on the monocultured fibroblasts and macrophages, I was curious to see how their expression profiles shifted in the context of co-culture. However, this is not a simple experiment, as there is always a concern that one cell type can ‘contaminate’ the other. I returned to this problem towards the end of my summer work, and conducted pilot tests and optimized protocols to purify and separate cells using magnetic beads.
Xu had previously noted that stimulation with high doses of Tumour Necrosis Factor alpha (TNFα) (100 µg/µL), an inflammatory cytokine, led to a qualitatively different response in cocultured cells compared with lower doses (10 µg/µL). These changes were not observed in monocultured cells. A graduate student in the lab, Josh, followed up with this observation using colonic stromal cells in place of the MEFs.
He used them to set up a colonic stromal cell and macrophage coculture, and I tagged along with MEFs. We worked together, alongside one another. However, during fluorescence-activated cell sorting (FACS) analysis, there were no macrophages detected in our experiments. Because it occurred in both sets of samples, we suspected it was the batch of macrophages that we used. While we weren’t able to get clean data from this particular experimental run, it was a useful experience to learn the considerations involved in this type of experimental setup, and to learn FACS analysis, a standard tool in cell biology and immunology (Figure 4). This was useful when I later worked on the bead separation protocol.
Module 3: Pilot LCMS for co-cultures
Complex, multicellular organisms have differentiated cells for the purpose of division of labor, including metabolism. Presumably, co-culturing different cell types that each have innately different metabolic capacities, would lead to a nonadditive shift in their overall metabolic capacity. LCMS seemed like a good first-pass approach toward answering this question.
In parallel to this wet lab work, I was trying to get up to speed on metabolism, in which I had less direct contact with experts in this area. As I researched different methods to deconvolute the metabolic profiles of two cell types grown together, a major problem seemed to be separation of the cells to individually evaluate their metabolisms. The very process of trypsinizing — or cutting proteins to release cells from a petri dish — is known to have a large impact on the metabolic profile of those cells. However, for the purpose of asking whether co-culturing leads to a non-additive effect to the metabolic profile, doing LCMS on the population of unseparated cells seemed adequate for an initial test.
I did a pilot test of preparing LCMS samples in my last week of the fellowship, but don’t yet have data.
Module 4: Engineered myddosomes
The Zhou lab has previously observed that stimulating cells with inflammatory cytokines in monoculture, compared with co-culture, leads to different responses. A cellular immune response leads to a whole cascade of effects, including alterations in metabolism, but that makes it difficult to discern the role and effects of individual players. More specifically, in response to a pathogen, immune cells form myddosomes, temporary organelles that transduce pathogen detection into an immune response.
Laura Landau from the Kagan lab previously created engineered myddosomes, a sort of temporary organelle that forms in the context of an immune response, functioning as the transducer between pathogen recognition and the cell’s immune response. Engineered myddosomes can be artificially stimulated to trigger a subset of the immune cascade (Figure 5). For example, TBK1 is an enzyme that contributes to the upregulation of glycolysis during an inflammatory response.
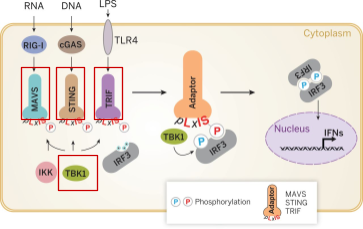
With Lauren’s help, I learned to transfect our cells of interest (CIM, a conditionally immortalized macrophage line, and 3T3, an immortalized fibroblast cell line) with her plasmids. Half of them failed (the CIM cells suffered contamination) and half of the 3T3s died off (likely plated too sparsely). But 3 plasmids (MAVS, STING, TRIF) are now safely tucked away in a -80C freezer. They can be later used to help interrogate how the various subsets of the immune cascade are relevant for mounting an effective immune response.
Conclusion
Summary: Many scientists thought that 3 months was too short of a time to do anything scientifically significant. They were right. Learning benchwork protocols from others is clearly the express route; it’s more efficient and comprehensive than reading protocols in papers. I found myself piggybacking on projects others around the lab were working on so that I could learn the relevant tools. Benchwork feels similar to programming, in that it's very useful to have a “reference” of code that runs, even if imperfectly, and subsequently adapt it for your needs.
Overall, this summer still enabled me to achieve my main goal; I learned about an area of interest, and got a foothold in experimental biology.
Future directions: I fortunately had the opportunity to stay another year, so the experience I’ve gained in the summer will hopefully mature and culminate into something more comprehensive, hopefully.
In the near term, I’d like to finish troubleshooting the cell type separation and build an RNAseq dataset on fibroblasts and macrophages grown in co-culture. Those experiments would then be expanded into monoculture vs coculture conditions with various inflammatory stimulations and, later on, I’ll examine the altered immune responses with engineered myddosomes. The next cycle of experiments will follow the resulting computational analyses.
In parallel, I would like to make more progress in investigating the metabolic attributes of the coculture system, starting with LCMS to get a rough idea of how the metabolic system changes, and followed by stable isotope tracing to get a more particular view of how the system processes metabolites.
In the long term, I aspire to be a ‘fullstack’ biologist, familiar with the full cycle of developing a hypothesis and model from reviews, to predictions and generating data to both test and perturb living cells.